The combination Design Engine for solid tumors.
We predict the target combinations that shut down escape routes in solid tumors, where 90% of cancer deaths occur.
We predict the target combinations that shut down escape routes in solid tumors, where 90% of cancer deaths occur.

Cell-line synergy metrics, HTS screens, and simplified model systems often miss the clinical context that determines patient benefit: resistance history, toxicity, line of therapy, biomarker context, and tumor evolution.
Most solid tumors adapt through redundant pathways, lineage plasticity, immune evasion, and resistance rewiring. Pharma needs a systematic way to identify combinations before committing to expensive trials.
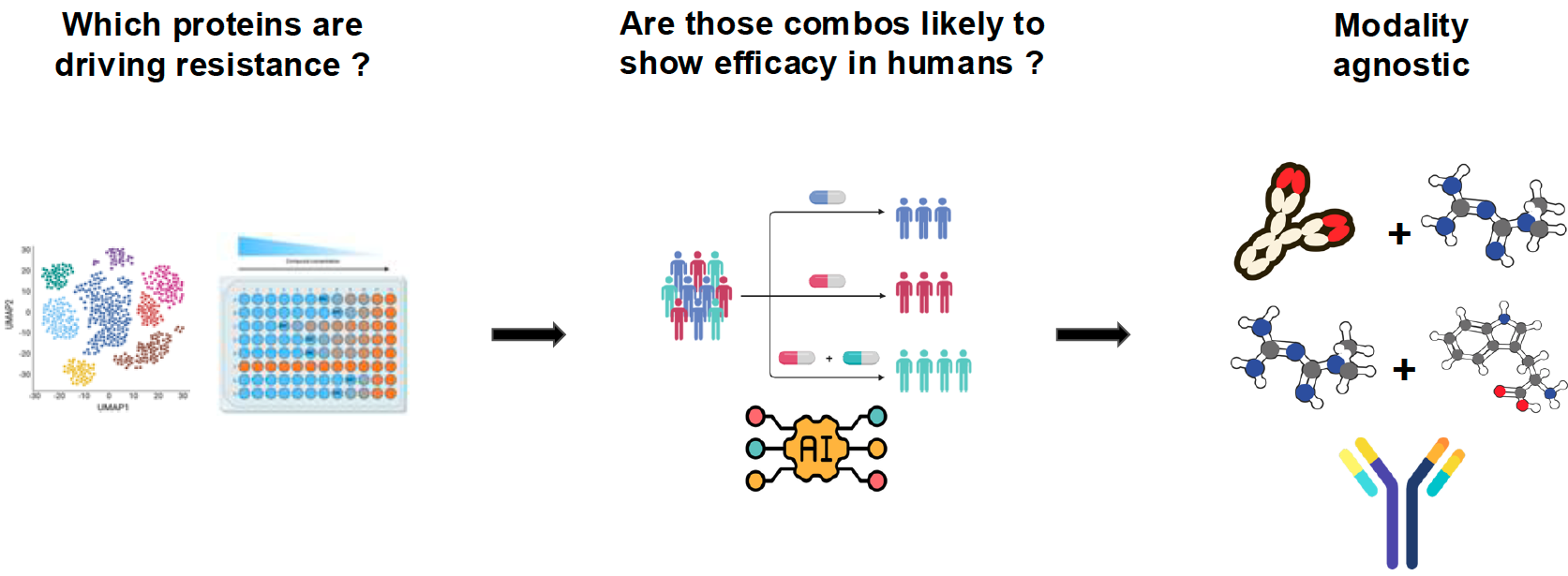
EMphora prioritizes human clinical outcomes as the primary translational signal while integrating mechanistic evidence from preclinical studies, literature, and target biology.

Mi earned his PhD at Heidelberg University under Prof. Julio Saez-Rodriguez in cancer systems biology, publishing in Nature Biotechnology and Cell Systems. He led the NCI-CPTAC DREAM Proteogenomics Challenge, coordinating 100+ scientists globally, then joined Stanford under Prof. Ash Alizadeh to build a computational reverse translation platform.




15+ years scaling biotech from pre-seed through IPO. Founding team member at Aligos Therapeutics, built all non-scientific functions, and led drafting of the S-1 business section ahead of the $150M IPO. Previously VP of Operations at Gordian Biotechnology. Earlier career in life sciences investment banking and strategy consulting.



Director of Oncology Clinical and Translational Bioinformatics at Moderna, applying machine learning to oncology drug development at scale. Previously Associate Director of Data Science at Novartis, leading exploratory biomarker analysis in clinical development. Built computational biology and biomarker platforms at H3 Biomedicine.




Head of Research at EMBL-EBI. Professor of Medical Bioinformatics at Heidelberg University and co-director of the DREAM Challenges. Postdoctoral training at Harvard Medical School and MIT. His group applies mathematical models and AI to large biological datasets to uncover the molecular basis of disease, with a focus on single-cell data, multi-omics integration, and cancer.





Cambridge, MA · celviontx.com